Mission
The Molecular Genomics Core (MGC) is a state-of-the-art multiomics facility equipped with high-sensitivity experimental platforms, enabling tailoring of experiments to meet researcher's genetic, genomic, epigenomic and proteomic needs. We work diligently to facilitate the success of your project from planning, data generation, and analysis.
Services
The MGC is proud to perform 10x Genomics assays, including Chromium scRNAseq/snRNAseq, Visium transcriptomic spatial gene expression, and Xenium in situ sequencing, alongside other genetic & proteomic experiments.
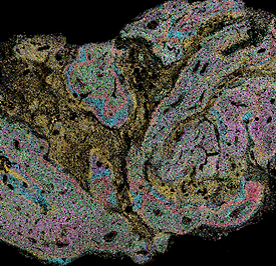
Header image is Xenium GBM data generated by the MGC for Simon Gregory & Lauren Whaley